Out-Of-Sample Predictions#
import arviz as az
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import pymc as pm
import seaborn as sns
from scipy.special import expit as inverse_logit
from sklearn.metrics import RocCurveDisplay, auc, roc_curve
RANDOM_SEED = 8927
rng = np.random.default_rng(RANDOM_SEED)
az.style.use("arviz-darkgrid")
Generate Sample Data#
We want to fit a logistic regression model where there is a multiplicative interaction between two numerical features.
# Number of data points
n = 250
# Create features
x1 = rng.normal(loc=0.0, scale=2.0, size=n)
x2 = rng.normal(loc=0.0, scale=2.0, size=n)
# Define target variable
intercept = -0.5
beta_x1 = 1
beta_x2 = -1
beta_interaction = 2
z = intercept + beta_x1 * x1 + beta_x2 * x2 + beta_interaction * x1 * x2
p = inverse_logit(z)
# note binomial with n=1 is equal to a Bernoulli
y = rng.binomial(n=1, p=p, size=n)
df = pd.DataFrame(dict(x1=x1, x2=x2, y=y))
df.head()
| x1 | x2 | y | |
|---|---|---|---|
| 0 | -0.445284 | 1.381325 | 0 |
| 1 | 2.651317 | 0.800736 | 1 |
| 2 | -1.141940 | -0.128204 | 0 |
| 3 | 1.336498 | -0.931965 | 0 |
| 4 | 2.290762 | 3.400222 | 1 |
Let us do some exploration of the data:
sns.pairplot(data=df, kind="scatter");
/home/xian/mambaforge/envs/pymc/lib/python3.10/site-packages/seaborn/axisgrid.py:118: UserWarning: This figure was using constrained_layout, but that is incompatible with subplots_adjust and/or tight_layout; disabling constrained_layout.
self._figure.tight_layout(*args, **kwargs)
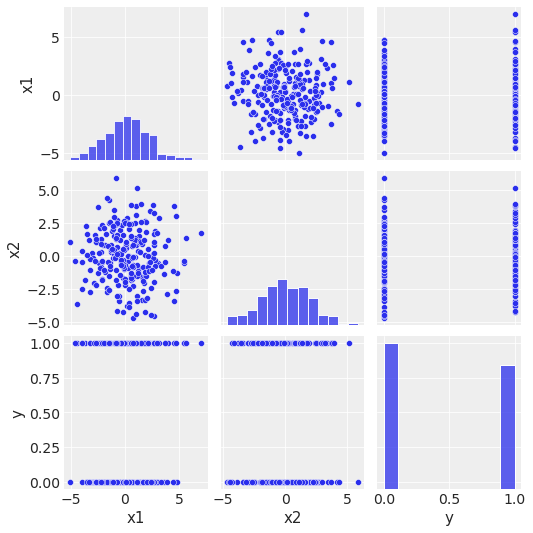
\(x_1\) and \(x_2\) are not correlated.
\(x_1\) and \(x_2\) do not seem to separate the \(y\)-classes independently.
The distribution of \(y\) is not highly unbalanced.
fig, ax = plt.subplots()
sns.scatterplot(x="x1", y="x2", data=df, hue="y")
ax.legend(title="y")
ax.set(title="Sample Data", xlim=(-9, 9), ylim=(-9, 9));
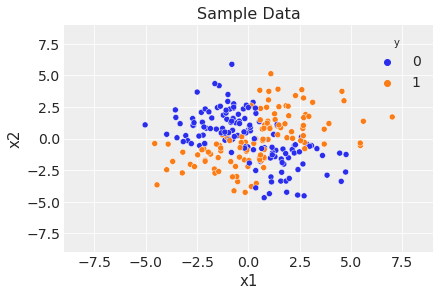
Prepare Data for Modeling#
labels = ["Intercept", "x1", "x2", "x1:x2"]
df["Intercept"] = np.ones(len(df))
df["x1:x2"] = df["x1"] * df["x2"]
# reorder columns to be in the same order as labels
df = df[labels]
x = df.to_numpy()
Now we do a train-test split.
indices = rng.permutation(x.shape[0])
train_prop = 0.7
train_size = int(train_prop * x.shape[0])
training_idx, test_idx = indices[:train_size], indices[train_size:]
x_train, x_test = x[training_idx, :], x[test_idx, :]
y_train, y_test = y[training_idx], y[test_idx]
Define and Fit the Model#
We now specify the model in PyMC.
coords = {"coeffs": labels}
with pm.Model(coords=coords) as model:
# data containers
X = pm.MutableData("X", x_train)
y = pm.MutableData("y", y_train)
# priors
b = pm.Normal("b", mu=0, sigma=1, dims="coeffs")
# linear model
mu = pm.math.dot(X, b)
# link function
p = pm.Deterministic("p", pm.math.invlogit(mu))
# likelihood
pm.Bernoulli("obs", p=p, observed=y)
pm.model_to_graphviz(model)
with model:
idata = pm.sample()
Auto-assigning NUTS sampler...
Initializing NUTS using jitter+adapt_diag...
Multiprocess sampling (4 chains in 4 jobs)
NUTS: [b]
Sampling 4 chains for 1_000 tune and 1_000 draw iterations (4_000 + 4_000 draws total) took 1 seconds.
az.plot_trace(idata, var_names="b", compact=False);
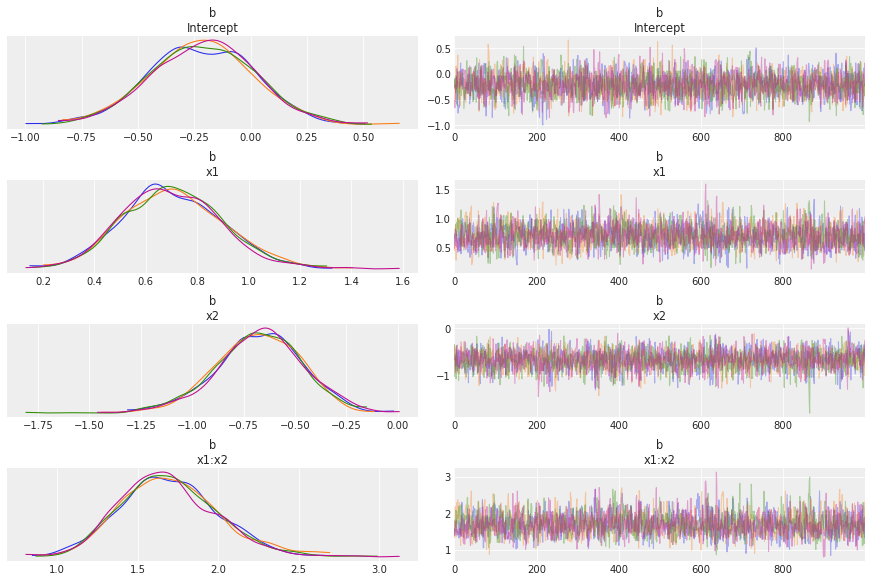
The chains look good.
az.summary(idata, var_names="b")
| mean | sd | hdi_3% | hdi_97% | mcse_mean | mcse_sd | ess_bulk | ess_tail | r_hat | |
|---|---|---|---|---|---|---|---|---|---|
| b[Intercept] | -0.215 | 0.231 | -0.650 | 0.216 | 0.004 | 0.003 | 3037.0 | 2565.0 | 1.0 |
| b[x1] | 0.705 | 0.193 | 0.363 | 1.085 | 0.004 | 0.003 | 2296.0 | 2386.0 | 1.0 |
| b[x2] | -0.675 | 0.209 | -1.065 | -0.281 | 0.004 | 0.003 | 2761.0 | 2419.0 | 1.0 |
| b[x1:x2] | 1.698 | 0.303 | 1.114 | 2.240 | 0.007 | 0.005 | 2079.0 | 2325.0 | 1.0 |
And we do a good job of recovering the true parameters for this simulated dataset.
az.plot_posterior(
idata, var_names=["b"], ref_val=[intercept, beta_x1, beta_x2, beta_interaction], figsize=(15, 4)
);
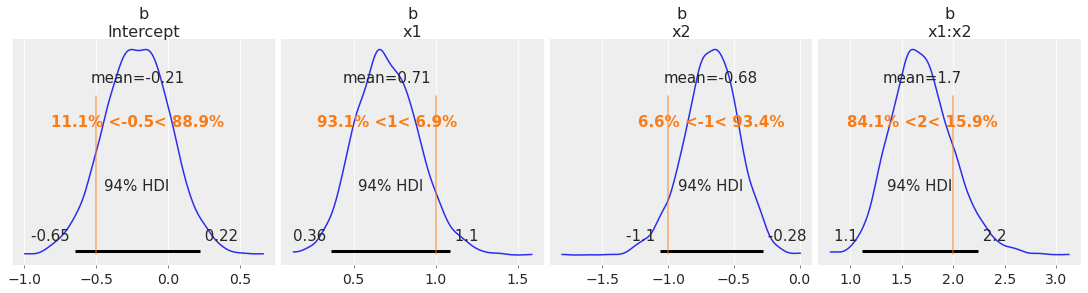
Generate Out-Of-Sample Predictions#
Now we generate predictions on the test set.
with model:
pm.set_data({"X": x_test, "y": y_test})
idata.extend(pm.sample_posterior_predictive(idata))
Sampling: [obs]
# Compute the point prediction by taking the mean and defining the category via a threshold.
p_test_pred = idata.posterior_predictive["obs"].mean(dim=["chain", "draw"])
y_test_pred = (p_test_pred >= 0.5).astype("int").to_numpy()
Evaluate Model#
First let us compute the accuracy on the test set.
print(f"accuracy = {np.mean(y_test==y_test_pred): 0.3f}")
accuracy = 0.893
Next, we plot the roc curve and compute the auc.
fpr, tpr, thresholds = roc_curve(
y_true=y_test, y_score=p_test_pred, pos_label=1, drop_intermediate=False
)
roc_auc = auc(fpr, tpr)
fig, ax = plt.subplots()
roc_display = RocCurveDisplay(fpr=fpr, tpr=tpr, roc_auc=roc_auc)
roc_display = roc_display.plot(ax=ax, marker="o", markersize=4)
ax.set(title="ROC");
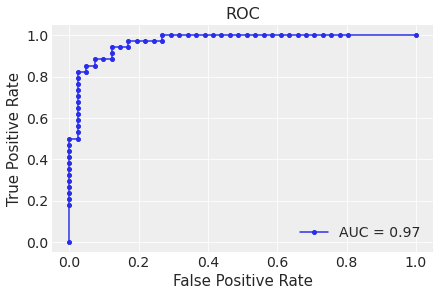
The model is performing as expected (we of course know the data generating process, which is almost never the case in practical applications).
Model Decision Boundary#
Finally we will describe and plot the model decision boundary, which is the space defined as
where \(p\) denotes the probability of belonging to the class \(y=1\) output by the model. To make this set explicit, we simply write the condition in terms of the model parametrization:
which implies
Solving for \(x_2\) we get the formula
Observe that this curve is a hyperbola centered at the singularity point \(x_1 = - \beta_2 / \beta_{12}\).
Let us now plot the model decision boundary using a grid:
def make_grid():
x1_grid = np.linspace(start=-9, stop=9, num=300)
x2_grid = x1_grid
x1_mesh, x2_mesh = np.meshgrid(x1_grid, x2_grid)
x_grid = np.stack(arrays=[x1_mesh.flatten(), x2_mesh.flatten()], axis=1)
return x1_grid, x2_grid, x_grid
x1_grid, x2_grid, x_grid = make_grid()
with model:
# Create features on the grid.
x_grid_ext = np.hstack(
(
np.ones((x_grid.shape[0], 1)),
x_grid,
(x_grid[:, 0] * x_grid[:, 1]).reshape(-1, 1),
)
)
# set the observed variables
pm.set_data({"X": x_grid_ext})
# calculate pushforward values of `p`
ppc_grid = pm.sample_posterior_predictive(idata, var_names=["p"])
Sampling: []
# grid of predictions
grid_df = pd.DataFrame(x_grid, columns=["x1", "x2"])
grid_df["p"] = ppc_grid.posterior_predictive.p.mean(dim=["chain", "draw"])
p_grid = grid_df.pivot(index="x2", columns="x1", values="p").to_numpy()
Now we compute the model decision boundary on the grid for visualization purposes.
def calc_decision_boundary(idata, x1_grid):
# posterior mean of coefficients
intercept = idata.posterior["b"].sel(coeffs="Intercept").mean().data
b1 = idata.posterior["b"].sel(coeffs="x1").mean().data
b2 = idata.posterior["b"].sel(coeffs="x2").mean().data
b1b2 = idata.posterior["b"].sel(coeffs="x1:x2").mean().data
# decision boundary equation
return -(intercept + b1 * x1_grid) / (b2 + b1b2 * x1_grid)
We finally get the plot and the predictions on the test set:
fig, ax = plt.subplots()
# data
sns.scatterplot(
x=x_test[:, 1].flatten(),
y=x_test[:, 2].flatten(),
hue=y_test,
ax=ax,
)
# decision boundary
ax.plot(x1_grid, calc_decision_boundary(idata, x1_grid), color="black", linestyle=":")
# grid of predictions
ax.contourf(x1_grid, x2_grid, p_grid, alpha=0.3)
ax.legend(title="y", loc="center left", bbox_to_anchor=(1, 0.5))
ax.set(title="Model Decision Boundary", xlim=(-9, 9), ylim=(-9, 9), xlabel="x1", ylabel="x2");
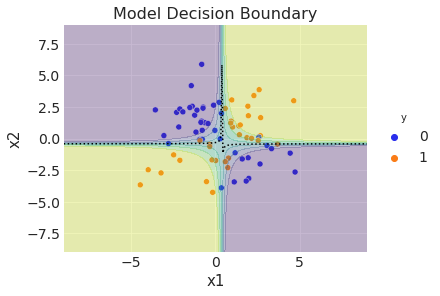
Note that we have computed the model decision boundary by using the mean of the posterior samples. However, we can generate a better (and more informative!) plot if we use the complete distribution (similarly for other metrics like accuracy and AUC).
References#
Watermark#
%load_ext watermark
%watermark -n -u -v -iv -w -p pytensor
Last updated: Wed Dec 06 2023
Python implementation: CPython
Python version : 3.10.8
IPython version : 8.4.0
pytensor: 2.18.1
pymc : 5.10.0
seaborn : 0.12.2
pandas : 1.4.3
matplotlib: 3.5.2
arviz : 0.14.0
numpy : 1.24.2
Watermark: 2.3.1
License notice#
All the notebooks in this example gallery are provided under the MIT License which allows modification, and redistribution for any use provided the copyright and license notices are preserved.
Citing PyMC examples#
To cite this notebook, use the DOI provided by Zenodo for the pymc-examples repository.
Important
Many notebooks are adapted from other sources: blogs, books… In such cases you should cite the original source as well.
Also remember to cite the relevant libraries used by your code.
Here is an citation template in bibtex:
@incollection{citekey,
author = "<notebook authors, see above>",
title = "<notebook title>",
editor = "PyMC Team",
booktitle = "PyMC examples",
doi = "10.5281/zenodo.5654871"
}
which once rendered could look like:
